Face masks, social distancing, lockdowns: Around the world, countries are racing to stop the transmission of the novel coronavirus SARS-CoV-2, and to mitigate the devastating effects of COVID-19, the disease it causes. Scientists, of course, have a major role to play in the global effort, modelling the epidemic and aiding in the development of new drugs.
But one group of scientists has been working on another, entirely different way of confronting viruses and preventing infection—editing the human genome. Launched in 2016, following a (somewhat secretive) meeting of more than 130 scientists, lawyers, and ethicists at Harvard Medical School, the Human Genome Project-write (HGP-write) set out to synthesize and edit human and other genomes. The project—which is named after the Human Genome Project, the effort completed in 2003 to sequence or “read” the entire human genome—originally intended to synthesize the full human genome—chemically combining the DNA building blocks to create synthetically what we all exist of naturally.
Weekly Newsletter
Several pilot projects were planned to develop the technology to get there, including creating a virus-resistant human cell line, editing the pig genome so that the animals could serve as human organ donors, and constructing chromosome versions as models for cancer. Over time, the price tag for DNA synthesis was expected to drop. However, in 2018, as the funding turned out to be scarcer than initially expected, the plan was downsized for the near future: The project’s first focus would be “recoding” the human genome, to produce cells immune to any kind of virus infection. This immunity would include known viruses as well as unknown viruses (like SARS-CoV-2 before 2020)—or, in short, all viruses.
By 2018, the project’s name had also been downsized—losing the “Human”—to just Genome Project-write, or GP-write. Despite the scaled-back scope, the goal is still ambitious: In the human genome, as well as in the genome of (nearly) all other living beings, each gene is written in a “four-letter alphabet,” with the letters corresponding to four chemical bases: adenine, thymine, guanine and cytosine (often referred to as A, T, G and C). Recoding would mean changing the way in which a cell uses and translates this genetic code, a process that has evolved over millions of years, with the goal of conferring viral immunity.
Optimizing Nature’s Alphabet
The genetic code contains apparent redundancies. Genes instruct the cell to select among 20 amino acids, which are the building blocks of proteins. How the cell strings together amino acids to form a functional protein is determined by triplets of bases, referred to as codons. The codon TCA, for example, stands for the amino acid serine. TCA tells a part of the cellular machinery, the tRNA, to grab serine and add it to the growing protein. Another codon, TAC, tells it to grab and add tyrosine. TAA, on the other hand, is a stop codon, and tells the cellular machinery to stop adding amino acids. But AGT also means serine, as do a number of other combinations.
GP-write proposes to optimize this code. Optimizing would mean that, out of all the codons that code for serine, only one gets picked—for instance, TCA. Any instances of redundant serine codons, like AGT, are then replaced with a chosen codon. This frees up the redundant codons to take over new functions. For instance, a freed-up codon could code for one of the hundreds of amino acids out there, rather than just the 20 that living beings currently string together into proteins. Organisms could produce novel enzymes and other proteins, unlike those made by living beings so far.
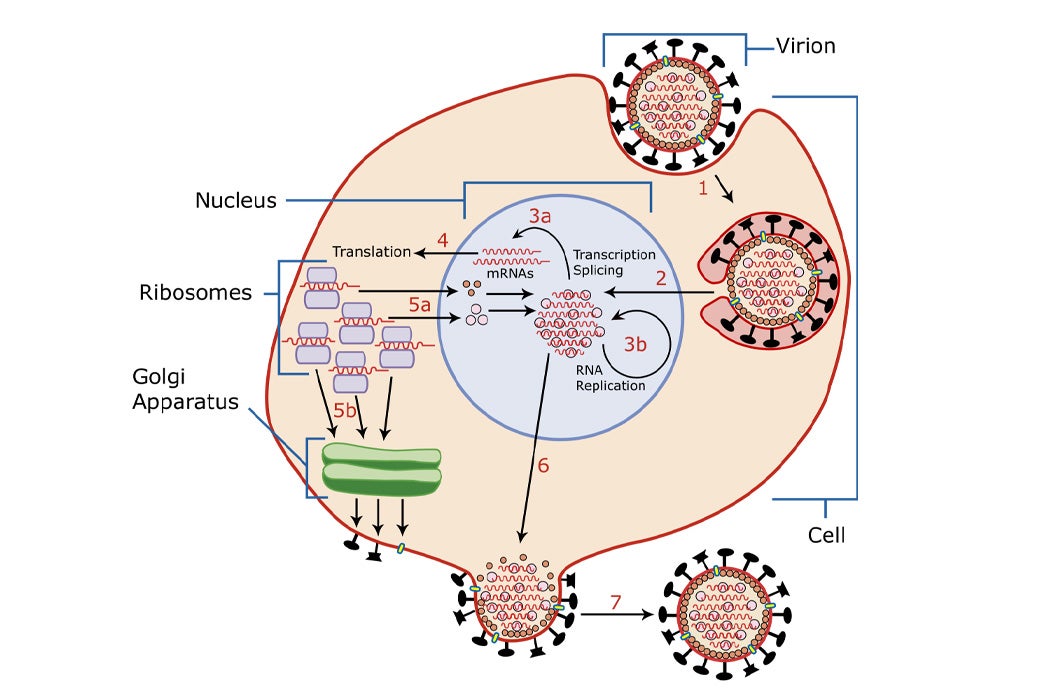
Re-assigning codons could make cells resistant to viruses. Normally, when a virus enters a cell, it hijacks the cell’s genetic machinery to produce more viruses, which are then released out into the world to infect others, through coughs, sneezes, and other means. In the recoded cell, the cellular machinery has been switched up: The tRNA that would grab serine in response to the (redundant) AGT codon is removed and replaced by a new tRNA, one that grabs a different amino acid when it encounters AGT. When a virus enters a recoded cell and attempts to reproduce through its cellular machinery, it simply can’t get the cell to produce more viruses. Either the cell just doesn’t recognize AGT as the instruction for serine anymore, or it adds a completely different amino acid in response to AGT—in both cases, the viral proteins that form are not functional. And so the infection stops.
Would this work not just for individual cells, but whole humans? Would a recoded human be resistant to all viruses, past, present, and future? George Church, one of the scientists spearheading GP-write (who, incidentally, is also interested in resurrecting extinct mammoths), has suggested so. In his 2012 book, Regenesis: How Synthetic Biology Will Reinvent Nature and Ourselves, he writes: “One of the greatest human health innovations of all time would be to make ourselves multivirus resistant… We could (also) achieve multivirus resistance by changing our genetic code.”
In theory, what works for a cell could, after further steps of optimization, work for an organism. So, when GP-write launched in a meeting that was closed to the media and the public, suspicion arose about the motives of the participating scientists.
The Ethics of Recoding
With the more focused aim of producing cells resistant to infection, announced in 2018, creating virus-resistant humans is off the table. GP-write clarified as much in a 2019 “Statement of Principle,” which states that “the consortium… will not use genome engineering technologies to create or modify human embryos.”
But the scientists go on to say: “Still, the eventual creation of genome engineering technology would in principle further enable the ability to engineer the genome of cells that are used to create and/or modify human embryos. Nevertheless, GP-write as a consortium has committed to a policy that it will not do so.” The statement was released after news broke that scientists affiliated with GP-write wanted to start a designer baby business, albeit one unconnected with GP-write itself.
Genome editing doesn’t just raise questions about designer babies. As the bioethicists Gregory Kaebnick, Michael Gusmano, and Thomas Murray put it in a 2014 report: “The threshold question for synthetic biology is whether the very prospect of synthesizing organisms is intrinsically or inherently problematic, whether or not it turns out to be good or bad for people.” However, they argue that, while synthetic biology is the “highly skilled alteration of nature,” it doesn’t necessarily change our relationship to nature, a question that should be resolved by public debate.
In 2010, after the first synthetic bacterium was produced, President Barack Obama requested a report on the ethics of synthetic biology. Amy Gutmann, Chair of the Presidential Commission for the Study of Bioethical Issues, recommended an approach of “prudent vigilance:” while it is possible that knowledge about creating synthetic organisms is put to malevolent “dual use,” the risks were seen as insufficient to limit experimenting with synthetic biology. “A responsible process will continue to evaluate safety and security as technologies develop. It will also include mechanisms for limiting their use when indicated,” Gutmann wrote in an essay.
The philosopher Mark Bedau argues that tinkering with the building blocks of life might, in itself, have scientific value:
The nature of life remains one of the deepest fundamental mysteries about our world. Once we finally figure out how to make wholly new forms of life entirely from nonliving materials, we will be able to design research programs to probe what kinds of radically novel chemical systems deserve to be considered as alive.
This research confronts us with the very basics of human nature: “The awareness of life’s arbitrary reprogrammability will no doubt shock and disturb some people. …That is, synthetic biology drives us to acknowledge that we, too, are complex chemical mechanisms.”
On the Shoulders of Bacteria
Even though GP-write’s sights have been set slightly lower, how achievable is its goal? Recoding the human genome will build on efforts in George Church’s lab to recode the genome of the bacterium Escherichia coli, making it resistant to viruses. In that project, Church and his team swapped out all 321 instances of the stop codon UAG, replacing it with UAA, also a stop codon. The researchers then deleted the gene that allows the cell to read the UAG codon. Nothing much changed for the recoded bacterium, which just grew more slowly. But a lot changed for viral invaders: as expected, they were no longer able to replicate well in the recoded host.
Extending this technique of removing and swapping codons to the much longer human genome won’t be easy. Replacing just a single codon in all 20,000 human genes takes hundreds of thousands of DNA changes. Another, potentially easier option is to make the genome from scratch, adding together letter after letter of the DNA, as the project initially proposed.
This research field, genome synthesis, has scaled up. It started with the first synthesis of the small hepatitis C genome, in 2000. From there, researchers went on to produce the first live organism with a working synthetic genome, a bacterium called Mycoplasma mycoides JCVI-syn1.0, in 2010. Next came the construction of several heavily rewritten chromosomes of baker’s yeast, Saccharomyces cerevisiae, a beloved eukaryotic model organism, in 2014 and 2017. Just last year, scientists succeeded in synthesizing the genome of E. coli and, in the process, replacing all occurrences of two codons for amino acids and one stop codon. Instead of the usual genetic code consisting of 64 codons, the bacterium follows a new set of rules with just 61 codons. And the recoded bacteria are alive: they grow more slowly, and they are longer and more rod-shaped than “standard” E. coli bacteria. But they are alive.
Synthesis could go even further: Synthetic biologists are trying to not just reuse codons, but to expand the DNA alphabet, adding in new letters in addition to the usual four. By adding in a new base pair, researchers could design semisynthetic organisms that build proteins from 172 different amino acids, quite an extension from the 20 amino acids currently used. Others seek to expand the genetic code by changing the basic rules, using quadruplet codons, e.g., AGGA instead of AGA, to incorporate unnatural amino acids.
If GP-write achieves its first goal, recoded cells may first be used in biotechnology. There, human cell lines are used to attenuate viruses for vaccines. The cell lines also serve as test beds to develop new therapies. Many companies use genetically engineered microbes to make medicines or chemicals. If a virus hits the cell lines, costly catastrophe can ensue.
Viruses can also be devastating to the dairy industry, which uses bacteria to ferment yogurt and cheese. The dairy industry sometimes has to dump its products when hit with viral contamination. Recoded organisms could also, some argue, be more “safe and useful” in such applications, as DNA released into the environment can no longer be picked up by natural (non-synthetic) organisms. Of course, the technique also can be put to nefarious uses, producing and engineering infectious viruses, the opposite of what GP-write’s scientists set out to achieve.
For practical and ethical reasons, making humans virus-resistant will have to wait. And genome recoding likely will not provide an answer to the coronavirus pandemic. But it opens up a tantalizing prospect: What if we stop playing by the same rulebook as viruses?